Salvatore Marras Laboratory
In the laboratory, which I share with Sanjay Tyagi and Fred Kramer at the Public Health Research Institute , a center within New Jersey Medical School at Rutgers University in Newark, New Jersey, I explore and develop self-quenching fluorescent nucleic acid hybridization probes and highly specific and sensitive nucleic acid amplification assays.
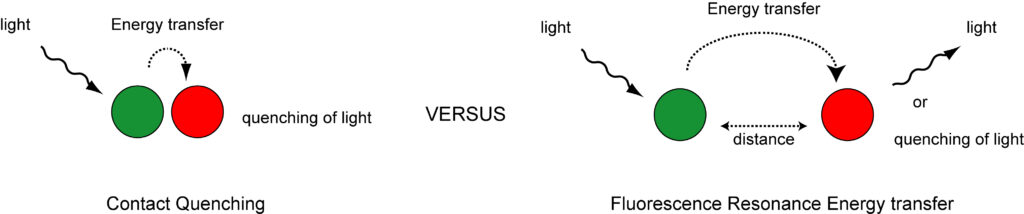
The introduction of self-quenching fluorescent nucleic acid hybridization probes has made it possible to detect RNA and DNA molecules in vitro, in vivo, in situ, and in silico with high sensitivity and specificity. My research focuses on the different properties of fluorescent nucleic acid hybridization probes. For example, I study the effect of the probe’s backbone chemistry (DNA, 2′-O-methyl-RNA, or LNA) on nucleic acid hybridization and I study the interactions between fluorophore and quencher pairs, which are being used as labels in these probes.
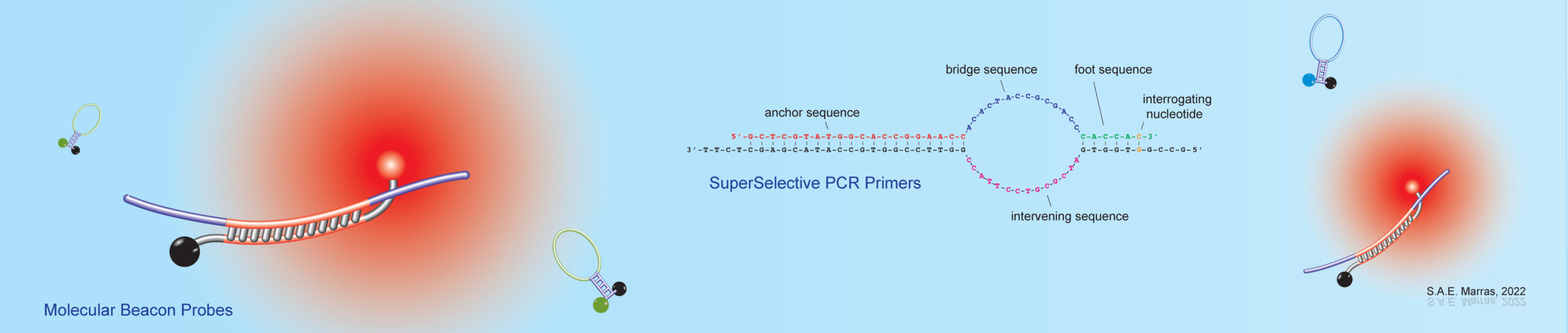
The work in our laboratory has lead to the development of the molecular beacon probe, one of the first fluorescent nucleic acid hybridization probe technologies. Molecular beacons have found their application in real-time monitoring of nucleic acid amplification assays, such as the Polymerase Chain Reaction (PCR), and isothermal amplication assays, such as Nucleic Acid Sequence Based Amplification (NASBA) and Loop Mediated Isothermal Amplification (LAMP) assays, which are being utilized for research and development and in clinical diagnostics. Real-time nucleic acid amplification assays provide both qualitative and quantitative information on rare RNA and DNA target sequences. Furthermore, these assays can be carried out in sealed tubes, thereby eliminating carryover contamination. Since molecular beacons remain dark when not hybridized to a nucleic acid target sequence, they also enable detection of DNA and RNA targets in living cells. You can find more information on molecular beacons, what they are, and what they can do, on the following web site www.molecular-beacons.org.
Novel nucleic acid detection methods and assays include highly multiplexed real-time nucleic acid amplification assays for clinical diagnostic detection of infectious agents, extremely sensitive, luminescent based in-situ and in-vivo hybridization methods, and self-reporting DNA micro array platforms.
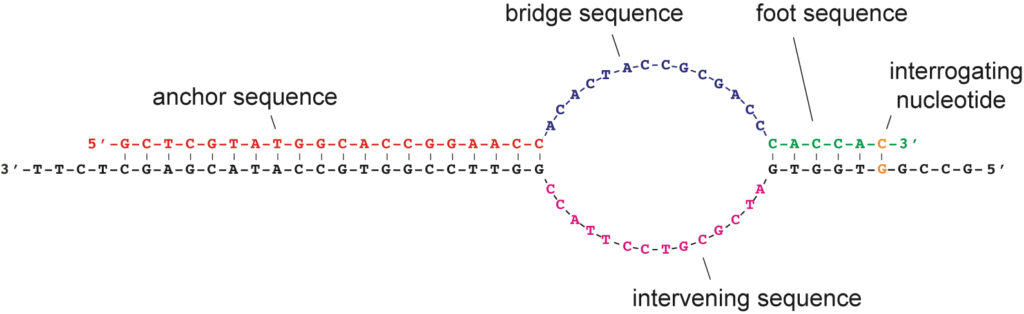
Recently, we introduced the SuperSelective primer technology, which enables the detection and identification of rare mutant DNA fragments in a large background of wild-type DNA fragments. We have shown their utility for the detection and quantitation of somatic mutations, whose presence relates to cancer diagnosis, prognosis, and therapy, in real-time multiplex PCR assays. SuperSelective primers can also be used to identify drug resistant and heteroresistant bacterial and fungal infections.